It's only taken 30 years, but information about Ebola in nature is finally starting to snowball. First, after almost 15 years of disappearing from the human population, Ebola returned with a vengeance in the mid 1990s, causing illness in 6 separate outbreaks in Gabon, Ivory Coast, Democratic Republic of Congo (DRC), and South Africa (imported case) between 1994 and 1996. As doctors and scientists rushed in to contain the outbreaks, they were also able to collect viral samples, and trap animals and insects in the area, searching for a reservoir for the virus. In this decade, there have been almost yearly outbreaks of Ebola and/or the closely related Marburg virus in Africa, resulting in the discovery of both Ebola and Marburg infection in species of fruit bats--suggesting these animals may be a reservoir species for filoviruses (though more work remains to be done to confirm this).
As I blogged about previously, prior work has suggested that the most deadly Ebola subtype, known as Ebola-Zaire (EBO-Z) after its initial site of isolation, has been spreading steadily eastward across the central African continent. This was tracked by examining isolates of the virus obtained during human epidemics, which introduces a bias into the sample. However, viral isolates from other sources have been quite difficult to obtain, despite many years of searching. A new paper examines viral isolates collected from dead gorillas and reconstructs their phylogeny in an effort to fill in some of these gaps; more after the jump.
Though human Ebola outbreaks get most of the attention, the virus has actually been far worse on other primates, particularly chimps and gorillas. Unfortunately though, we've mostly only seen the carnage in these primate populations after the fact, when workers at wildlife sanctuaries stopped seeing apes they'd become familiar with, or researchers happened upon their corpses in the forest.
The latest paper used some of these tragic deaths to further our understanding about Ebola. Over a 5 year time period, researchers found 47 dead animals in the Gabon/Republic of Congo region--17 of these were determined to be infected with Ebola-Zaire. Using the polymerase chain reaction, they were able to amplify portions of the glycoprotein gene (GP) from 6 gorillas and a chimpanzee, and compare the sequences to those previously identified in humans. When they compared the sequence to other EBO-Z GP sequences published to date, they found that these new genes were divergent enough to constitute a new group within the EBO-Z subtype--and that recent human cases from the Republic of Congo during the same time frame also fit into this new group (designated group B; the previous identified groups were group A, which included sequences from the 1976 outbreak and several outbreaks in the mid-1990s, and group R, from outbreaks between 2001-3).
They also sequenced a second viral gene, encoding the nucleoprotein (NP). For this they were only able to get sequence from 2 gorillas, as well as from human outbreaks during this time period. They found again that the sequences from this gene were closely related to each other, but more divergent from previously deposited sequences. Interestingly, some of the viruses that fit into group A by virtue of their GP sequence weren't found to be genetically distinct from the group B viruses when looking at the NP sequence, which the authors suggest could be due to a recombination event.
They next track the recent outbreaks across Africa by EBO-Z group (green: oldest lineages, group A; blue, group R; red, group B.)
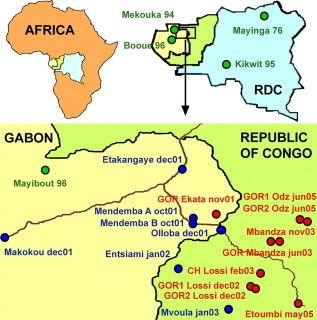
While upon initial glance this appears to support the previous research regarding Ebola's eastward spread, the authors argue against this simpler explanation, noting that their new data show that the most recent outbreaks weren't caused by descendants of previously emerging viruses, but by genetically different types of EBO-Z. Therefore, this draws into question the previous research painting EBO-Z emergence as the radiation of a single viral lineage (acquiring minor mutations along the way), and instead suggests the spread is due to multiple introductions of related viruses (perhaps moved along by a reservoir species?)
They also note that even for corpses found in the same place, infections from several independent sources may have occurred. For example, they describe a 2002 find of 2 gorillas in the Lossi park, which were found next to each other and likely died around the same time. However, when they examined the GP gene sequences, they were found to differ at 11 bases, suggesting "independent evolution for a minimum of 8 months" based on their molecular clock analysis. Extending this analysis to all viruses, they hypothesize a common ancestor of all Ebola examined in this study existed only ~30 years ago--or right around the time of the first identified human outbreak in the Democratic Republic of Congo (then Zaire).
Though this new research is interesting, it once again highlights the glaring gaps in our knowledge. They conclude:
What remains unclear is how often such introductions have occurred and to what extent transmission among susceptible animal hosts, such as great apes, may have subsequently contributed to the spatial propagation of outbreaks. Similarly, it remains to be resolved whether the temporal-spatial patterns of EBO-Z emergence, which are in many ways suggestive of a spreading process, could be the result of transmission processes in the reservoir species or whether other factors could have generated such patterns.
This research and the previous transmission paper they build on are starting to plug some holes, as did the discovery of the virus in fruit bats, long thought to be potential reservoirs or carriers of Ebola. It's still baby steps for now, but at least it's progress toward a better understanding of this still-mysterious pathogen.
Reference
![]() TJ Wittmann et al. 2007. Isolates of Zaire ebolavirus from wild apes reveal genetic lineage and recombinants. PNAS. 104:17123-27.
TJ Wittmann et al. 2007. Isolates of Zaire ebolavirus from wild apes reveal genetic lineage and recombinants. PNAS. 104:17123-27.
Image from http://www.pnas.org/content/vol104/issue43/images/large/zpq041077933000…
- Log in to post comments

Great post. Thanks.
Well, now there are two gaps, right?
Do the anti-germ theory gang have anything to say about Ebola? I'm wondering if they (a) pretend it doesn't exist; (b) claim there's some other, natural, completely non-viral reason why people start turning into chunky soup; or (c) claim that Ebola outbreaks are actually the CIA testing their secret orbital microwave people-liquifying technology, and virus researchers are covering it up.
Do the anti-germ theory gang have anything to say about Ebola?
Of course. It's the same nonsense, or non-science, your choice, as HIV, marburg, H5N1, west-nile, coronavirus, you name it.
Not new viruses are the bad news. Dumb virologists, not aware they're watching ancient phenomena with new spectacles, like PCR, all excited they've discovered something unheard-of, together with chemists who pretend poisoning is good health-care and World Hype Organizations preaching about one new global panic per month, there's your bad news.
Wow. That made even less sense than most of the things that jspreen says.
What, exactly, jspreen, do you think causes the symptoms of Ebola? Or do you deny that these outbreaks actually occur?
What, exactly, jspreen, do you think causes the symptoms of Ebola? Or do you deny that these outbreaks actually occur?
Why should I deny things that really seem to happen? I don't deny that people are ill, I just think that the self proclaimed health experts of today simply know nothing about why people are ill and have completely lost all track of the slightest idea of what life is all about.
Give me a grant and I'll spend a couple of month where it all happens, speak with the people up there, and I'll tell you what exactly causes Ebola.
I vaguely remember a great phrase of Kary Mullis I read somewhere ages ago, something about a pandemic caused by an Ebola virus that couldn't even make it to the next village, but I can't find the pages where I read it. But Google came up with an interpretation of Ebola which I find infinitely more interesting than the We-fight-viruses-with-killer-chemo non-science.
Here's an extract of the article:
Ebola and marburg are viruses that attack our endothelial cells and cause bleeding but if you look closely at the literature it's the prolonged release of Nitric Oxide from our white blood cells causing most of the damage. We just get burnt by our own acid.
I was watching a show on Ebola one day and was horrified as the highly paid virus warriors in their protective white suits just put a Congolese lady into a corner with a bowl of water and watched her die.
What did they expect to happen?
So I want you to think carefully about what we are doing here, it's actually complete negligence based on our paranoid belief system of pathogens.
I believe that Ebola is only occurring in areas of low Selenium and Iodine combined with low protein diets and scurvy.
We could save people with a truckload of Brazil nuts, Lysine tablets, Lemons and maybe a few cans of Tuna or Salmon and a dash of iodized salt.
jspreen ,
Do you have a map with zone of low selenium ? so we can see if there is overlap with the ebola outbreak or no.
This would allow to check if you have a (small) point or no.
Selenium is important in the glutathione cycle and as an anti-viral per si.
Reduced immune system defense due to lack of selenium and low plasma glutathione, and therefore suceptibility to Ebola dont invalidate the fact that Ebola may be caused by a virus.....
Hey, no fair asking jspreen for evidence of anything. He said it, he believes it, that settles it :)
Reduced immune system defense due to lack of selenium and low plasma glutathione, and therefore suceptibility to Ebola dont invalidate the fact that Ebola may be caused by a virus.....
Yours is of course a perfectly true reasoning and it helps us understand why the We-fight-viruses-with-killer-chemo non-scientists always perfectly get their asses away from the fire each time their nonsense threatens to blow up in their face.
Okay, jspreen, since you're such a heroic crusader, off you go to the Congo with your brazil nuts and tuna fish. Once you've cured everyone, you can come back and tell us all about it, 'kay?
Okay, jspreen, since you're such a heroic crusader, off you go to the Congo with your brazil nuts and tuna fish. Once you've cured everyone, you can come back and tell us all about it, 'kay?
They're not my brazil nuts and tuna fish, but the author of the cited text's, but I find your offer extremely generous. After I've put some money I borrowed in plane tickets, nuts and fish, I feed the poor who have previously been stolen by the rich, where after, in case I succeeded to make them feel better even though the robbery goes on, you allow me to come back. Grand!
(Emphasis mine)
Could anyone please tell me the average annual income, including salary, bonuses or any other money which goes to the personal spending of a "virus warrior"? Of all the ones I have met (which is very few to be honest), calling them "highly paid" would make welfare close to nobility.
calling them "highly paid" would make welfare close to nobility.
Again you don't get it at all, apy. Compared to poor and malnourished, dying from ebola Congolese ladies, virus warriors are rich as Cresus.
Jan, you may note that many of the people helping to fight outbreaks such as Ebola are those who live in the outbreak-stricken areas. I'm sure you didn't read the recent National Geographic article on zoonotic diseases that I linked (it's here), but several of the featured "virus warriors" themselves had family that died from Ebola.
You should also note that Ebola has stricken gorillas in Lossi gorilla sanctuary as I note--and these gorillas ain't lacking for food.
What is your point jspreen? Compared to Congolese ladies, a teenagers allowance is "highly paid". Everyone reading this blog is "highly paid", even you, if that is the comparison they are making. Why is the article saying that if the point is not to some how claim the "virus warriors" are making huge profits off spreading claimed fear and misinformation? Would "virus warriors" only be accepted if they worked for free? Or could not afford to feed themselves?
jspreen, surely you wouldn't want me to offer you any money at all to perform your no doubt vital and lifesaving brazil-nut delivery run. If I did, you would be a highly-paid brazil nut warrior, and we would be entitled to reject anything you said on the subject, as you'd obviously be biased. You're going to have to prove that you are not in any way beholden to the powerful brazil nut lobby if you want to have any credibility here.
"Give me a grant and I'll spend a couple of month where it all happens, speak with the people up there, and I'll tell you what exactly causes Ebola."
Can I just say that sending jspeen to study Ebola up close and personal sounds good to me.
I think we may have stumbled on a key point here. Unless and until jspreen provides conclusive proof to the contrary, we have to assume that he's a shill for the nutritional-supplement industry, which makes a fortune selling unproved, untested, often useless and sometimes harmful products to the ignorant and unwary. Nothing he says can be trusted. After all, compared to a starving Congolese villager, he must be as rich as Bill Gates.
Sending jspreen to study Ebola up close may be an attractive offer to some but it is doubtful to ever happen, a grant would make jspreen highly paid, then we'd be reading about how both too little money and too much money cause disease. Deciding how to spend all that twenty thousand dollar salary that viral warriors make can be very strenuous.
jspreen, surely you wouldn't want me to offer you any money at all to perform your no doubt vital and lifesaving brazil-nut delivery run.
Why should I want money for that? Where do I enter the scene? Anybody can send the nuts or whatever directly to the people who need it.
Nothing he says can be trusted.
Right you are. Nothing. I'm after burning pants and to judge the average writings of the Ebolavirus-apologists, I'm doing quite well.
According to what metric?
So:
jspreen thinks that Ebola can be cured with Brazil nuts;
and that he could find the "real" cause of Ebola in a couple of months;
but he doesn't have any intention of actually sending any Brazil nuts to the Congo;
or of going there himself.
Conclusion: all mouth, no trousers. Burning or otherwise.
I'll put up a bag of brazil nuts and $100 towards a one-way airfare for Spreen to go to the DRC. Anyone else want to contribute?
I'll put in $100. Although there has to be some verifiable way to make sure jspreen is working in the DRC.
Man, the best grant/funding suscription ever! Hey, this is a historical moment. Come on folks, let's come together and make this work. I'll put in $100 from my own pocket. But, and I hope you didn't forget, a one-way ticket won't do. I must be able to come back and report, according to Wells' initial proposal. And eventually we'll think of some other more or less minor details to work out, but let's sit back and see what happens for the next couple of days, alright?
Maybe we can get a DonorsChoose fund going. Education in the classroom/DRC!
"Give me a grant and I'll spend a couple of month where it all happens, speak with the people up there, and I'll tell you what exactly causes Ebola."
You mean Ebola isn't caused by bad emotions one or two months before diagnosis, which was preceded by good emotions? Damn, I thought you had it all figured out.
An interesting Ebola blog
http://www.msf.ca/blogs/ZoeY.php
Yes, interesting blog, really. I read only half of it but I think I already know what Ebola is mostly all about. Just another deadly stroke of fear-mongering Center of Dummy Control and World Hype Organization officials. It's strange, the blog-writer writes it down herself but she doesn't seem to realize what she writes.
Meanwhile, a child had also come in with a fever and a history of contact. She was so sad sitting there bored and probably pretty scared with all of us wandering around in our suits.
Yeah, you got it. And they rub it in. Look at the suits and the masks and all. Be assured, it will give any healthy person the creeps. I wonder how this man felt when addressed by a such a thouroughly protected health care specialist. If I were him, I'd have felt like some piece of deadly vermin and I would have instantly dropped dead in mortal shame.
Be sure, my friends, if ever our grant subscription gets me to the DRC, I'll never wear nor mask nor suit. But then again, who would accept such a fool amidst the healthy and the sick?
Seems y'all don't discuss the original topics of entries much. Usually bogs down to name calling but if that is what you guys like to do....
And the CIA doesn't have the melty ray satellite anymore. Bill Gates bought it to cook popcorn.
Man, I thought the anti-academic trolls were bad at Inside Higher Ed, but this thread sets some kind of record for allowing a troll to drive the discussion. Congratulations, I guess.
A troll that doesn't drive the discussion is a pretty poor excuse for a troll. Most every one I've observed gets well over 50% of the comments in the thread directed at himself. Sometimes on topic comments are so rare as to be a notable exception.
Now the psychology of the troll and his victims, that's an interesting subject. OT of course.
It would seem that with renewed outbreaks, progress can now be made. How unfortunate that more will have to die so that sufficient data can be collected until ultimately a cure can be found.
If nutritional deficiencies were the sole cause, the epidemiological patterns would not be anywhere near what has been observed, in this layman's opinion.
If nutritional deficiencies were the sole cause, the epidemiological patterns would not be anywhere near what has been observed
The epidemiological pattern here is quite comparable to the epidemiological pattern of H5N1 bird flu. Every when and where you put the right people with the right instructions to look for what you want them to find, they'll dig up the thing.
I suppose the between the lines part here is that it is obvious H5N1 bird flu is a result of nutritional deficiencies?
I suppose the between the lines part here is that it is obvious H5N1 bird flu is a result of nutritional deficiencies?
Leave the between the lines for the more experienced reader, apy, and limit yourself to correctly decoding of the printed words. In case you'd still need some further explanation after having read what's written black on white: H5N1 as the cause of bird flu, let alone mutating H5N1 as the cause of some human variant of bird flu pandemic, is bullshit nobody can top.
Uhh so you're saying my between the lines statement was correct? Thanks for the condescending post only to tell me I was right.