tags: Bovine spongiform encephalopathy, BSE, mad cow disease, Creutzfeldt-Jakob Disease, CJD, pathogenic mutation, prion protein gene
Mad Cow Disease, technically known as Bovine Spongiform Encephalopathy (BSE), is one of a group of transmissible diseases that destroy brain tissue, collectively known as Transmissible Spongiform Encephalopathies (TSEs). TSEs are an unknown agent(s) that act by damaging the structure of brain proteins known as "prions" (PREE ons). In turn, these damaged prion proteins damage other normal prions and together, they build up to collectively destroy tissue in the brain stem, causing cavities to develop in the victim's brain so it resembles swiss cheese in mammals. This neurodegenerative disease rapidly leads to death for the victims, of course.
In humans, this disease has long been known as Creutzfeldt-Jakob Disease (CJD) in honor of the two medical scientists who first described it in the literature. But is the agent that causes BSE in cattle the same one that causes CJD in humans? What are TSEs; genetic diseases, infectious agents, or sporatic mutations? Where did the first TSE come from? These basic questions about the nature of TSEs have remained unanswered for decades -- until today, that is.
Today, a paper was released by the open-access journal, PLoS Pathogens, that was written by two veterinary scientists, Jürgen A. Richt and S. Mark Hall, from the United States Department of Agriculture. These scientists show that a rare pathogenic mutation underlies BSE that is identical to CJD, and further, their study also supports the hypothesis that BSE originated from a previously undetected TSE from contaminated cattle feed in the UK.
To do this work, the researchers isolated proteins from the brainstems of a variety of animals and humans that had been diagnosed with a TSE. They performed immunohistochemical studies on these proteins and confirmed the presence of an abnormal Prion Protein, PrPd, that characterizes Mad Cow Disease and other TSEs (figure 1);
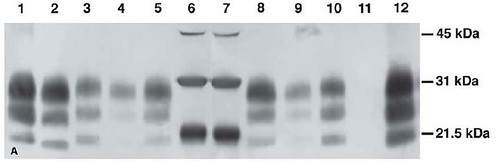
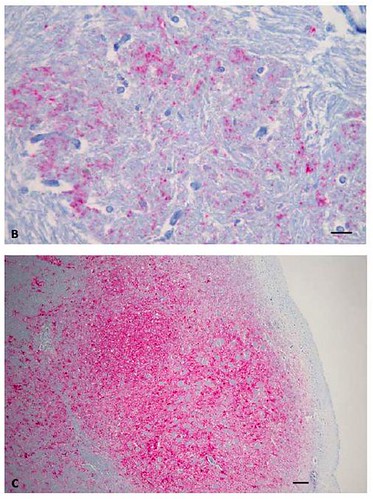
Figure 1. Analysis of brainstem samples from BSE-infected animals employing various methods. (A) Hybrid Immunoblot Analysis using enriched samples: Lanes 1-5: monoclonal antibody 6H4 (raised against human PrP residues 144-152), lanes 8-12 monoclonal antibody P4 (raised against ovine PrP residues 89-104): 1 = sheep scrapie control, 2 mg; 2 = classical BSE (2003 U.S. BSE case), 2 mg; 3 = H-type BSE case (2004 U.S. BSE case), 2 mg; 4 = U.S. BSE Alabama case, 1 mg; 5 = U.S. BSE Alabama case, 2.5 mg; 6,7 = protein weight maker; 8 = U.S. BSE Alabama case, 2.5 mg; 9 = U.S. BSE Alabama case, 1 mg; 10 = H-type BSE case (2004 U.S. BSE case), 2 mg; 11 = classical BSE (2003 U.S. BSE case), 2 mg; 12 = sheep scrapie control, 2 mg. [larger view] (B) Immunohistochemistry of the U.S. BSE Alabama case (H-type BSE) using PrP-specific monoclonal antibody F99/97.6.1. Brainstem at the level of obex was examined. Bar = 35 µm. (C) Immunohistochemistry of a classical BSE case [4] using PrP-specific monoclonal antibody F99/97.6.1. Brainstem at the level of obex was examined. Spongiform changes are found in the area with highly PrPd-positive cells. Bar = 90 µm. [larger view].
doi:10.1371/journal.ppat.1000156.g001
In the above figure, 1A provided information regarding the molecular size and relative amount of this protein, while figures 1B and C revealed the location of these abnormal proteins in the brainstem tissues at death.
After confirming the presence of the abnormal proteins, the researchers then isolated DNA from each animal and sequenced the prion protein genes. To distinguish minor differences between these genes, they aligned all the DNA sequences next to each other and compared mutations in each (Figure 2);
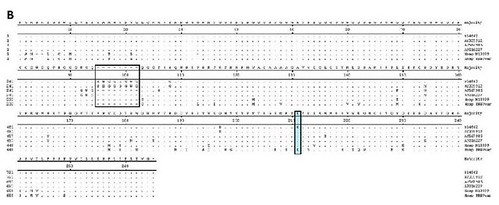
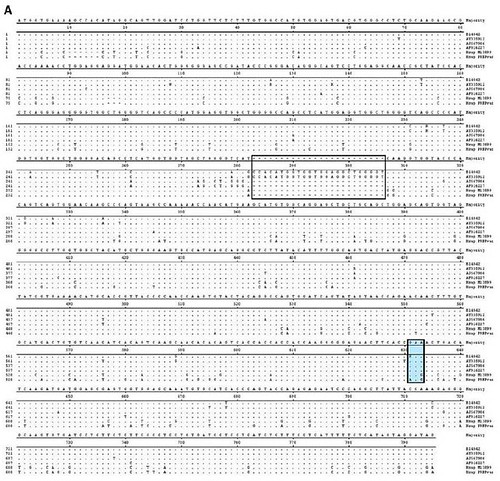
Figure 2. Alignment of bovine, ovine, cervid and human Prnp sequences. (A) Nucleotide sequences. Standard single letter codes are used for nucleotides. Y = C or T; R = A or G; K =G or T; W= A or T. Boxed area indicates the 6th octapeptide-repeat of the bovine protein (U.S. BSE Alabama case and sequence AY335912). Additional Prnp sequences are as follows: AJ567986 (sheep), AF016227 (elk), Hsap M13899 (human, normal) and Hsap PRNPvar [human, variant; see [13]]. [larger view]. (B) Amino acid sequences. Standard IUPAC single letter codes are used for amino acids. Codon numbering refers to the most common six-copy octapeptide repeat allele for Bos Taurus. Boxed area indicates the 6th octapeptide repeat of the bovine protein [animals B14842 [4] and AY335912]. AJ567986 (sheep), AF016227 (elk), Hsap M13899 (human, normal) and Hsap PRNPvar [human, variant; see [13]] each contain a 5 octapeptide repeat region in the protein. [larger view].
doi:10.1371/journal.ppat.1000156.g002
As a result of their work, the authors identified a novel mutation in the bovine prion protein, which they named E211K. As you can easily see in the above figure, this mutation is identical to the E200K pathogenic mutation in the human prion protein -- the most common cause of genetic CJD in humans.
The study lends support the hypothesis that all three forms of TSEs in humans are also found in cattle: infectious, sporadic, and genetic. It provides additional support that the BSE epidemic may have originated from a genetic case of BSE in an individual cow. Of course, similar mutations can pop up in cattle herds throughout the world and might provide the raw genetic material for new epidemics to develop in the future, especially in countries where it is a common practice to feed slaughterhouse scraps to healthy animals that will enter the human food chain in the future.
Source
Jürgen A. Richt, S. Mark Hall, David Westaway (2008). BSE Case Associated with Prion Protein Gene Mutation PLoS Pathogens, 4 (9) [PDF] DOI: 10.1371/journal.ppat.1000156.
- Log in to post comments



Interesting! The question that immediately came to mind was, "Can the abnormal PrP direct improper folding of normal PrP?" That would seem to be a requirement for the variant protein to cause disease transmissible to the general population.
Would a common lineage explain this similarity?
Assuming that this PrP^Sc originating in genetic TSEs is indeed the infectious agent responsible for infectious TSEs, would this explain the ease with which the infectious agent jumps the species barrier?
You seem to be implying here that ingesting PrP^Sc (originally) sourced from animals with genetic TSEs is indeed the cause of Infectious TSEs. Could you clarify?
If such is indeed the case, can we also conclude that sporadic TSEs are due to the same genetic defect occurring as a somatic mutation which creates a small amount of seed PrP^Sc to start the disease off?
Prof. Bleen -- excellent question and yes, indeed, the mutant PrP can and does refold the normal PrP protein, and those proteins then refold other normal PrP proteins, and they all build up until there are few, if any normally folded PrP left. this "behavior" of TSEs is what really shook the biochemical/microbiology world since it went against the established paradigms of the time: disease is caused by a microbe or by a faulty gene (not by an improperly folded protein).
Stagyar zil Doggo -- this mutation doesn't "jump the species barrier" in the typical sense of the word. the genetic form of this disease can be due to a spontaneously occurring mutation in the PrP gene, which is a protein that is found in the brains of all mammals. when this mutation occurs in reproductive (germ) cells, it can then be passed on to all that animal's offspring. incidentally, the epidemic that i refer to is the one that occurred in the UK during the 1980s and 1990s, which devastated their beef industry.
besides inheritance, this disease can be transmitted by eating the flesh of animals that were infected (as is often the case for cattle that are intended for slaughter). those animals, in turn, were infected either by eating infected flesh, or by a mutation, either a somatic or germ cell mutation.
i hope this answers your questions.
Hm, I remember reading about this in my high school bio textbook. I got the distinct impression that they were the same disease. I must have not been reading carefully enough, or the textbook was publishing a hypothesis like a conclusion.
Oh, and by the way, I think you will find it's pronounced "PRY-ons". [/snooty]
"Oh, and by the way, I think you will find it's pronounced "PRY-ons". [/snooty]"
Not preferably--sometimes given as a second correct pronunciation, though.
Thanks for this blog post GrrlScientist. By default or design, the field of TSE's is my main area of interest:-
http://network.nature.com/profile/steelgraham
Jürgen and Mark's PLoS Pathogens Manuscript came to my attention on Friday and I'm glad that it's getting some deserved attention.
I've had contact with Jürgen in the past and I again applaud USDA scientists for making their work Public Access.
--
It was a pleasure meeting you at sciblog and as I said, I was truly touched by the fact that readers of your blog made it possible for you to attend (and present at) the event in person.
"PRY on" is the preferred pronunciation when referring to the birds, prions -- six similar species of Procellariidae. at least, that's how i learned it from my microbiology and ornithology professors.
McDawg -- i was pleased to meet you, too! and i am glad you pointed out your area of expertise, but did you write about this paper yet? i looked at your blog but didn't see anything.
The content of my personal blog is evolving Grrl. Rather than blogging exclusively about Prion related issues, I'm pretty much decided that I'll try to stick to Neuroscience with a bit of silliness thrown in as well.
I have a teleconf on Thursday with the CJD International Support Alliance (CJDISA) and I'll make sure this Manu. is discussed. I've already emailed the CJDISA a copy of it.
The content of my personal blog is evolving Grrl. Rather than blogging exclusively about Prion related issues, I'm pretty much decided that I'll try to stick to Neuroscience with a bit of silliness thrown in as well.
I have a teleconf on Thursday with the CJD International Support Alliance (CJDISA) and I'll make sure this Manu. is discussed. I've already emailed the CJDISA a copy of it.
Sorry, I appear to have been unclear in my previous comment.
What I meant to ask was whether the said similarity in mutations in Bovine and Human genes is responsible for the fact that misfolded Bovine Prion Proteins can misfold Human Prion Proteins on contact (and presumably vice versa) resulting in Humans contracting vCJD by ingesting cows affected by BSE. Contrast this with TSEs in other animals like Scrapie or Wasting Disease which don't appear to be (so easily?) transmissible to humans.
are we to hepatitisX yet???
Thanks for submitting this post to our blog carnival. We just published the 39th edition of Brain Blogging and your article was featured!
Thank you.
Sincerely,
Shaheen