 Few things can take me out of blogging hibernation (especially when the next grant deadline is Monday...) However, one of those things that I'll carve out time to write about is an interesting, hot-off-the-presses Ebola paper, and especially one describing a new strain of the virus--and there just happens to be such a paper in the new edition of PLoS Pathogens. Details after the jump...
Few things can take me out of blogging hibernation (especially when the next grant deadline is Monday...) However, one of those things that I'll carve out time to write about is an interesting, hot-off-the-presses Ebola paper, and especially one describing a new strain of the virus--and there just happens to be such a paper in the new edition of PLoS Pathogens. Details after the jump...
Previously, 4 types of Ebola viruses had been identified. Ebola Zaire has typically been the worst as far as fatality rates (around 80-90%), with Ebola Sudan coming in close behind (roughly 50-70%). The other two types are both more rare and, to date, far less lethal. Ebola Reston was discovered in monkeys imported into the U.S. for biomedical research, and hasn't caused any known disease in humans (although several workers exposed to the infected animals did develop an antibody response to Ebola Reston, showing they had apparently been subclinically infected). Only one case of infection with the Ivory Coast strain of Ebola has been found to date: a scientist who had done a necropsy on an Ebola-infected chimpanzee later became ill. This was in 1994, and represents the last time a new Ebola subtype has been described.
However, there were rumblings of this new species of Ebola last year when I blogged about an Ebola outbreak in Uganda. This outbreak likely started in late August or early September of 2007, and was declared over in February 2008 after causing 149 suspected cases and 37 deaths (25% mortality, although the figure of 36% is given in the paper due to ongoing studies).
However, when researchers stepped in to investigate this outbreak, they noticed something funny. Antibody tests were coming back positive, but the initial tests looking for virus using a sensitive real-time PCR assay were negative. Using a less specific PCR assay, they carried out some preliminary work looking at a different area of the virus (the L gene), and found that its sequence didn't match any of the known Ebola types. To confirm this, investigators used "pyrosequencing" to determine the nucleotide sequence for more of the genome--a rapid method that gave them most of the viral genome in just a few days. From the draft genome, they were able to develop new primers specific for the novel virus strain (designated "Bundibugyo Ebolavirus") and therefore, have a new PCR test specific for this emergent Ebola strain.
They also compared Bundibugyo to other Ebola subtypes, and found that it was most closely related to the Ivory Coast species. This is interesting, as the area of Uganda where this outbreak took place was near the DRC border, and Uganda borders Sudan as well--and yet the Bundibugyo subtype was fairly distantly related to the (geographically more proximal) Sudan and Zaire strains. This despite the site of isolation of the Cote d'Ivorie strain being, by my very crude estimation, about 2400 miles away from Bundibugyo--just showing again how little we know about Ebola virus ecology. This brings up all sorts of unanswered questions--what other Ebola strains are circulating in Africa? Will we eventually find all of the current subtypes (and more new ones?) in fruit bats, as we have for Ebola Zaire and the other filovirus, Marburg? Why does the Bundibugyo type seem to cause less mortality in humans than the Zaire and Sudan strains? Would we see a similar phenomenon if the Cote d'Ivorie strain were to cause a large human outbreak? As always with filovirus research, this new paper brings up more new questions than it answers.
Jonathan S. Towner, Tara K. Sealy, Marina L. Khristova, César G. Albariño, Sean Conlan, Serena A. Reeder, Phenix-Lan Quan, W. Ian Lipkin, Robert Downing, Jordan W. Tappero, Samuel Okware, Julius Lutwama, Barnabas Bakamutumaho, John Kayiwa, James A. Comer, Pierre E. Rollin, Thomas G. Ksiazek, Stuart T. Nichol (2008). Newly Discovered Ebola Virus Associated with Hemorrhagic Fever Outbreak in Uganda PLoS Pathogens, 4 (11) DOI: 10.1371/journal.ppat.1000212
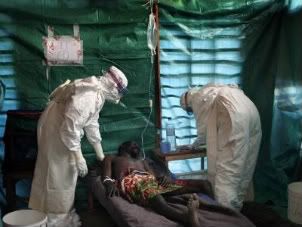
Image from http://www.javno.com/slike/slike_3/r1/g2008/m02/y164330786683117.jpg

Link is live - you can send a trackback now if you want.
Interesting.
Anyway, how many sentences on average does each of the 18 authors contribute in the referenced paper? ^_^ I know... just kidding.
Professor Smith, thank you VERY much for this. I had mentioned the report briefly in our undergraduate virology class yesterday but lacked such a concise and clear interpretation of the paper. If it's okay with you, I'm going to post the link on Blackboard for our class.
Good luck with that grant deadline!
Sure, post away.
Thanks for this Tara. We miss you.
I had mistakenly placed this comment on one of your above links to a former blog about ebola. I meant to place it here. Sorry.
I am not so sure about this ebola and marburg stuff as being only about a virus. I'm not saying that there is not a virus called ebola or marburg, and I am not saying that such a virus cannot be a, or even "the" major contributive factor, but there are other factors involved in Uganda, Congo, Sudan, etc, that are being completely overlooked and ignored.
For instance, the vast majority, almost all of the ebola/marburg cases turned up in Africa, were at or very near to gold mining camps, where cyanides, as well as various acids such as hydrochloric and sulfuric and acid, and aqua-regia are used, and often combined with cyanides resulting in an even more toxic brew. These well known toxic chemicals are almost always being used to get the gold from the ore. Some of the cyanide is potassium cyanide, and there are other cyanides including sodium and combined with hydrochloric acid, creating hydrogen cyanide, that are created or used as well. These are used in a process wherein the ore is crushed and soaked in cyanide. The gold dissolves into the cyanide solution and is then precipitated out. The cyanide mixture is often further treated with acids to purify it or burned.
Over in Africa, there is no control over what is done with the cyanides and acids and fume from combined solutions, and the spent solutions are usually dumped right onto the ground or into streams, and undoubtedly seep into ground water going to wells.
And when combined or burned, they often produce even more deadly toxic substances.
Cyanide is a potentially deadly poison. It works by making your body unable to use oxygen, without which life cannot be sustained. Certainly exposure to cyanide can weaken an individual to the point where other viral factors can take over to finish the job. Cyanide poisoning can happen quickly from single dose, or slowly from gradual repetitive lower exposures.
Furthermore, the majority of those affected by ebola/marburg, besides being in the toxic and polluted gold mining areas, are nearly all very poor, and the poverty stricken, and quite often even have insufficient diets, and surely also are further weakened by such contributing factors.
As such, in lower doses, these highly toxic chemicals can also slowly cause the deterioration of bodily function, thereby easily allowing other, normally nonharmful viruses, etc, to finish off the individual.
Many in Africa are unwittingly exposed to cyanides and acids used in mining, and do not know it, and they are never tested for such. Toxicologists are not even to be found there.
There is no known research that has ever been done on what the combined effects of some of these indigenous viruses are capable of when combined with factors such as cyanides, acids, fumes, and poor diets, so we really do not know what the truth of the matter is.
Such factors probably never will be known as long as the majority of aetiologists and virologists only consider viral causes as the sole root cause of these illness. It is unfortunate that there is very little crossover between toxicologists and virologists, nutritionists, etc, to look at the combined factors that are so obviously intertwined and certainly play their roles. Seldom do these differing specialized individuals even communicate with each other, let alone collaborate.
Specialization may be necessary in todays labor market, but it certainly is harmful in furthering science and getting to the facts of the matter of various human illness, and in discovering the intercourse of roles played by both pathogens and toxins, not to mention sufficient nutrition on any live creature including mankind.
AnotherTake:
Cyanosis does not cause fever or internal bleeding. Also, most large mining companies are careful to recycle their cyanide, as it is expensive to produce and transport.
Informal 'artisan' gold miners generally don't use cyanide for gold extraction; they use metallic mercury, as it is cheaper, easier to transport, and doesn't kill you straight away. While this causes severe health effects, those effects are different to both cyanide poisoning and viral infection. Also, Uganda's main gold mining areas are in the SE, near lake Victoria and Tanzania, while the ebola outbreak was on the Western border with the DRC.
Mr. Lab Lemming, first of all, my discussion was not that cyanide causes the symptoms, as I was referring to people being weakened by toxic and even nutritional factors, thereby allowing indigenous microbes to do their further damage, and the lack of collaborating studies on such, let alone consideration of such by virologists. Or perhaps even the effects of such toxins on the microbes in question.
Secondly, is there something that you know that the rest of us do not, about the effects of combining of various toxins such as cyanide and cyanic and acid fumes, not to mention mercuric fumes, and their relationship to the possibility of enhanced effects of various indigenous viraemia such as ebola or marburg and other indigenous pathogens in those already weakened by exposure to such various toxins, including the mercury fumes you mention, as well as by exposures to other toxins we have not yet mentioned, as well as combining these factors with those weakened by insufficient nutrition? Just what is it you know about these combined effects or the lack of effects that the rest of us are not privy to?
And if you know do know about these combined effects or lack of such effects of toxins, viruses, bacteria, and poor nutrition, it would be very nice if you would inform the rest of the world, since there are no collaborative scientific studies on the matter.
Thirdly, gold is found throughout Uganda by artisan miners, including on both sides of the western and even the southwestern borders, though you are correct that the larger industrial mines are in the SE (and some larger industrial gold mines are also in the SW, by the way).
Furthermore, many Ugandans have even recently been caught mining on the DR Congo side right across, and right next to the border with Uganda, including Ugandan military officers, and smuggling gold and ore back to Uganda. By the way, Mr. Lab Lemming, pray do tell.....: Which way does the wind blow and which way does the water flow from the artisan gold mines, discussed at the following bbc report, that are very very near to the border of Congo and Western Uganda?
http://news.bbc.co.uk/1/hi/world/africa/3879589.stm
As such, I must say that I find your response to my post just a bit off track, and, dare I say, even misleading by your suggesting there are no gold mines on the border, or anywhere near the ebola outbreaks.
You said: "while the ebola outbreak was on the Western border with the DRC".
Such was the exact point of the post I had written, Mr. Lemming. The artisan gold mining is on and near both sides of the border and near the outbreaks, and this being the exact point..... So I must Thank You for further confirming my suspicions, Mr. Lemming.
And I do like your name, Mr. Lab Lemming. Seems somehow appropriate to your, ummm, very thoughtful, response, and the point of my post. After all, lemming suicide is a frequently-used metaphor in reference to people who go along unquestioningly with popular opinion, with potentially dangerous or fatal consequences. This is also the theme of the video game "Lemmings", where the player attempts to save the mindlessly marching lemming rodents from walking to their deaths.
But do forgive me, Mr. Lab Lemming, as I certainly do not mean to stop you from marching to your own death. So, by all means, don't let me detain you, and do keep walking.
But during your death march, Mr. Lemming,
perhaps you could take a moment to look at this week old BBC report of a village of starving people on the eastern border of Congo and Uganda, and consider the effects of bodies weakened by starvation and consider what more could possibly be needed to allow for the indigenous viruses to finish them off:
http://news.bbc.co.uk/2/hi/africa/7725997.stm
To Another Take on it:
I am confused, are you saying that the filioviruses are not the causes of the diseases associated with them (ebola, marburg)? I understand that most of the people who become infected with ebola are in poor health due to their living conditions, but there is one very large exception.
Marburg virus was discovered when lab technicians in Marburg, Germany became ill after working with infected monkey tissue in 1967 (See Wikipedia). I believe that we can safely assume that these people were not suffering from the same malnutrition, parasitic loads, or environmental toxins as the people in Uganda and surrounding areas.
So while I think that we can agree that outbreaks in the Ugandan area are worse due to the population's health status, it is also quite clear that the filioviruses are the single causative agent of Ebola.
(As a side note, I don't believe that there is a great deal of argument about ebola being anything other than an indigenous virus. Perhaps I am missing your precise meaning of "indigenous'?)
Another:
I'm not an expert in this area, but your argument doesn't stack up to me on too many counts for me too consider it seriously. I'm not going to give all the objections (too many), but a few for thought:
As Lab Lemming pointed out, the symptoms of cyanide poisoning and Ebola infection are different. If cyanide poisoning preceded Ebola, cyanide poisoning symptoms would have been noted in early-stage cases, rather than those associated with Ebola. That seems contradictory to me.
Cyanide poisoning is considerably faster-acting than Ebola, killing within hours rather than days, and has no incubation period as Ebola, etc. do. How could it be cyanide poisoning if that would kill the affected people well before Ebola would?
How would experienced mine staff not recognise cyanide poisoning?
And before you say "only mild poisoning": you'd have to have every person in an Ebola outbreak "conveniently" have only mild poisoning, and have the mild poisoning make them more susceptible to viral infections despite cyanide poisoning being known for a long time and have other things that "weaken the body" induce specifically Ebola, and have to explain away Ebola cases in otherwise healthy people. Too many concidences, I think ;-)
And why suddenly be true now and only for Ebola/Marburg?
Likewise, vice versa: why would other incidents of cyanide poisoning in Africa in similar situations not be linked to infections?
Cyanide in various forms has been used for a long time in many industries: if there was a link with infection, it'd be well-known by now I would think.
From what I understand, potassium cyanide breaks down in the environment, so that doesn't seem very likely route for absorbing (much) of it. (The "environmental" argument would make more sense for mercury.)
Either way, what you suggest seems full of contradictions. I have to say it reads closer to cooler's style of rambling "alternative" causes for a disease than anything that I would consider seriously, certainly not without a much, much more substantial case for it.
http://blog.researchgate.net/index.php?/archives/36-New-Ebola-subtype-d…
You medics amuse me!
You actually seem to care more for science than wealth. No wonder Ebola still exists!
Actually chronic low level exposure to cyanide is well known to cause the exact same symtoms as Ebola and related viruses. The fevers and the internal bleeding and external bleeding from all orafices.
I remember reading a study a while ago about some otters at a zoo that were eating cyanide containing seeds. The low levels of cyanide in the seeds was not enough to cause acute cyanide poisening, however over time two of the otters came down with all the symtoms of hemorraghic fever and died of it. The study concluded that the cyanide was the cause of these symtoms.
People who come down with hemorraghic fevers are always exposed to cynaide and other potent toxins (just like the lab techs in Germany). Hemorraghic fevers travel down the river from mining town, just as the cyanide that is dumped in the river dose.
Most outbreaks of Ebola and related fever occurs when illeagal miners take over a closed mine. And yes they do use their own cyanide to extract the gold. Actually they often build cyanide pits in the very homes where their wives and children sleep. The miners get the Ebola the most followed by their wives, followed by their children. The rest of the village gets it at a lower rate.
Also in mining towns that come down with Hemorraghic fever the number of virus strains in the polulation matches the number of ethnic groups in the population. In other words the number of distinct genetic groups of people determine the number of virus strains.
This clearly shows that these viruses are endogenous creations of the sick person and not an infectious organism. This should come as no suprise since all viruses look and behave exactly like endogneous nucleic acid physiology and look and behave nothing like infectious organisms. When a nucleic acid is transcibed in a nonlinear way it is called a virus (dyanamic genome theory of viruses), when it is transcribed in a linear manner it is admited to be an endogenous nucleic acid.
All virological diseases follow this pattern, in humans, animals, plants, and bacteria. There is always a toxin or toxins that cause the exact same disease as the virus. The people, animals, or plants that come down with the viral disease are always exposed to these toxins. The toxin is the intial cause of the disease cause it causes the over production of the endogenouse viral sequence in the organism. The virus, which is really an endogenous cellular regulator, then causes the diese by virtue of it's over abundance.
A good example of this is the recenlty discovered obesity viruses. Yes, being fat is a viral disease. Yet obesity is not contagious. What happens is when a person over eats their body overproduces obeisty viruses and they put on weight. The obesity viruses, which is a type of adeno-virus, is a DNA virus that regulate stem cell development. It causes stem cells to develop into mature fat cells. It uses it's viral genes to guide the cell along it's development into a mature fat cell.
The obesity viruses are produced via non-linear transciption so as would be predicted modern scientist has once again misidentified them as infectious organisms and have put forth the absurd theory that you can catch fatness from other people. They have even recomened that hand washing is a good way to stay thin.
Daniel,
Clearly you need to take a look back to the last hundred+ years of biological discoveries. Even the simplest experiment with lambda phages in e. coli will show you that these are no endogenous sequences native to the host organism. Yes, humans, and most complex living organisms, carry endogenous retroviral elements. These however are most often transcriptionally silent due to the accumulation of mutations. However, if you wish to claim that Ebola is nothing more than an endogenous part of the human chromosome perhaps you will provide some evidence to back this up. Can you show where it is in the fully sequenced human genome? Maybe a paper or two identifying it's sequence in our chromosomes? Likewise, if you wish to claim that cyanide poisoning was responsible for the outbreaks of hemorrhagic fevers blamed on Ebola then surely you have real evidence for this. Perhaps blood tests from the victims, showing low levels of cyanide, or cyanide-derivatives? Please feel free to post links to said results as I'm sure you aren't just coming up with this BS without real, solid evidence...right?
Also, in regards to:
"Also in mining towns that come down with Hemorraghic fever the number of virus strains in the polulation matches the number of ethnic groups in the population. In other words the number of distinct genetic groups of people determine the number of virus strains"
Perhaps you can explain why your hypothesis would fail in light of the fact that the Marburg virus appeared both in Germany (where it was first identified after infecting German lab workers)and in local African populations.
If your hypothesis were correct, one would have to deduce that the African Margurg victims are more closely related genetically to the German lab workers (who shared a common virus) than those Africans infected with a different type of Ebola.
Do you truly think such a relationship is logical?
If you state the fact that Ebola is possibly nothing more than an endogenous part of the human chromosome should really be present facts.
we wonder about the places of ebola. only short african town are visited from the virus - i think, this is a test
I think you should take the virus seriously. One should, however, Africa is among the poorest countries so.