blastp
Sometimes when you go digging through the databases, you find unexpected things.
When I was researching the previous posts on insulin structure and insulin evolution, I found something curious indeed.
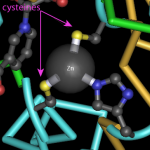
Human insulin, colored by rainbow. Image from the Molecule World iPad app by Digital World Biology.
I wanted to find out how many different organisms made insulin, so I used a database at the NCBI called Blink. Blink is a database of protein blast search results. Using Blink can save you lots of time because…
In which we identify unknown human proteins.
Yesterday, I wrote about using the BLOSUM 62 matrix to calculate a score for matches between two proteins. Those scores give us a good start on understanding how blastp determines whether two sequences are matching by chance or because they're more likely to be related. But that's not all there is to calculating a blast score, and there's at least one other statistic to consider as well, the E value.
It all comes down to biochemistry
The BLOSUM 62 matrix is based on the substitutions that really do or do not happen in real protein sequences. I…
In which we search for Elvis, using blastp, and find out how old we would have to be to see Elvis in a Las Vegas club.
Introduction
Once you're acquainted with proteins, amino acids, and the kinds of bonds that hold proteins together, we can talk about using this information to evaluate the similarity between protein sequences. We can easily imagine that if two protein sequences are identical, then those proteins would have the same kind of activity. But what about proteins that are similar in some regions, and not others, or proteins that only share some of the same amino acids in similar…